# Symbiodinium sp diversity in two coral species at two reefs (banks)
data(green.data)
# removing outliers
goods=purgeOutliers(
data=green.data,
count.columns=c(4:length(green.data[1,])),
zero.cut=0.25 # remove this line for real analysis
)## [1] "samples with counts below z-score -2.5 :"
## [1] "EFAV153"
## [1] "zscores:"
## EFAV153
## -3.884085
## [1] "OTUs passing frequency cutoff 0.001 : 10"
## [1] "OTUs with counts in 0.25 of samples:"
##
## FALSE TRUE
## 3 7# stacking the data table
gs=otuStack(
data=goods,
count.columns=c(4:length(goods[1,])),
condition.columns=c(1:3)
)
# fitting the model
mm=mcmc.otu(
fixed="bank+species+bank:species",
data=gs,
nitt=3000,burnin=2000 # remove this line for real analysis!
)## $PRIOR
## $PRIOR$R
## $PRIOR$R$V
## [,1] [,2] [,3] [,4] [,5] [,6] [,7] [,8]
## [1,] 1 0 0 0 0 0 0 0
## [2,] 0 1 0 0 0 0 0 0
## [3,] 0 0 1 0 0 0 0 0
## [4,] 0 0 0 1 0 0 0 0
## [5,] 0 0 0 0 1 0 0 0
## [6,] 0 0 0 0 0 1 0 0
## [7,] 0 0 0 0 0 0 1 0
## [8,] 0 0 0 0 0 0 0 1
##
## $PRIOR$R$nu
## [1] 7.02
##
##
## $PRIOR$G
## $PRIOR$G$G1
## $PRIOR$G$G1$V
## [1] 1
##
## $PRIOR$G$G1$nu
## [1] 0
##
##
##
##
## $FIXED
## [1] "count~otu+bank+species+bank:species+otu:bank+otu:species+otu:bank:species"
##
## $RANDOM
## [1] "~sample"
##
##
## MCMC iteration = 0
##
## Acceptance ratio for liability set 1 = 0.000175
##
## Acceptance ratio for liability set 2 = 0.000386
##
## Acceptance ratio for liability set 3 = 0.000246
##
## Acceptance ratio for liability set 4 = 0.000351
##
## Acceptance ratio for liability set 5 = 0.000351
##
## Acceptance ratio for liability set 6 = 0.000579
##
## Acceptance ratio for liability set 7 = 0.000439
##
## Acceptance ratio for liability set 8 = 0.000754
##
## MCMC iteration = 1000
##
## Acceptance ratio for liability set 1 = 0.209912
##
## Acceptance ratio for liability set 2 = 0.422000
##
## Acceptance ratio for liability set 3 = 0.218404
##
## Acceptance ratio for liability set 4 = 0.436123
##
## Acceptance ratio for liability set 5 = 0.352246
##
## Acceptance ratio for liability set 6 = 0.420579
##
## Acceptance ratio for liability set 7 = 0.360211
##
## Acceptance ratio for liability set 8 = 0.514632
##
## MCMC iteration = 2000
##
## Acceptance ratio for liability set 1 = 0.295000
##
## Acceptance ratio for liability set 2 = 0.426947
##
## Acceptance ratio for liability set 3 = 0.301140
##
## Acceptance ratio for liability set 4 = 0.451456
##
## Acceptance ratio for liability set 5 = 0.375421
##
## Acceptance ratio for liability set 6 = 0.418737
##
## Acceptance ratio for liability set 7 = 0.374018
##
## Acceptance ratio for liability set 8 = 0.526667
##
## MCMC iteration = 3000
##
## Acceptance ratio for liability set 1 = 0.306614
##
## Acceptance ratio for liability set 2 = 0.455596
##
## Acceptance ratio for liability set 3 = 0.320193
##
## Acceptance ratio for liability set 4 = 0.403772
##
## Acceptance ratio for liability set 5 = 0.391298
##
## Acceptance ratio for liability set 6 = 0.427105
##
## Acceptance ratio for liability set 7 = 0.418737
##
## Acceptance ratio for liability set 8 = 0.529070# selecting the OTUs that were modeled reliably
acpass=otuByAutocorr(mm,gs)
# calculating effect sizes and p-values:
ss=OTUsummary(mm,gs,summ.plot=FALSE)
# correcting for mutliple comparisons (FDR)
ss=padjustOTU(ss)
# getting significatly changing OTUs (FDR<0.05)
sigs=signifOTU(ss)
# plotting them
ss2=OTUsummary(mm,gs,otus=sigs)## ymax not defined: adjusting position using y instead
## ymax not defined: adjusting position using y instead
## ymax not defined: adjusting position using y instead
## ymax not defined: adjusting position using y instead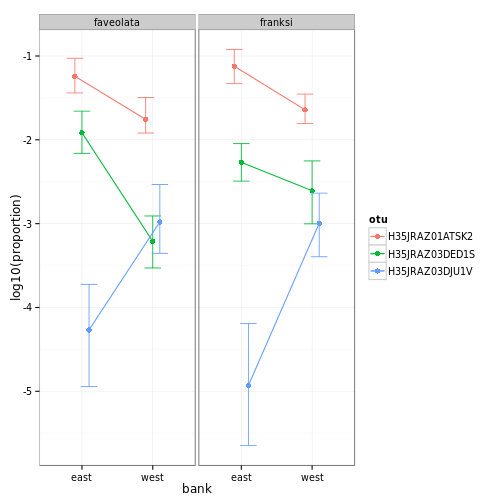
# bar-whiskers graph of relative changes:
# ssr=OTUsummary(mm,gs,otus=signifOTU(ss),relative=TRUE)
# displaying effect sizes and p-values for significant OTUs
ss$otuWise[sigs]## $H35JRAZ01ATSK2
## difference
## pvalue bankeast:speciesfaveolata
## bankeast:speciesfaveolata NA
## bankeast:speciesfranksi 0.76715746
## bankwest:speciesfaveolata 0.01826772
## bankwest:speciesfranksi 0.09050840
## difference
## pvalue bankeast:speciesfranksi
## bankeast:speciesfaveolata 0.120384169
## bankeast:speciesfranksi NA
## bankwest:speciesfaveolata 0.003861344
## bankwest:speciesfranksi 0.012605292
## difference
## pvalue bankwest:speciesfaveolata
## bankeast:speciesfaveolata -0.5142604
## bankeast:speciesfranksi -0.6346446
## bankwest:speciesfaveolata NA
## bankwest:speciesfranksi 0.7572422
## difference
## pvalue bankwest:speciesfranksi
## bankeast:speciesfaveolata -0.4002223
## bankeast:speciesfranksi -0.5206065
## bankwest:speciesfaveolata 0.1140381
## bankwest:speciesfranksi NA
##
## $H35JRAZ03DED1S
## difference
## pvalue bankeast:speciesfaveolata
## bankeast:speciesfaveolata NA
## bankeast:speciesfranksi 0.2747204819
## bankwest:speciesfaveolata 0.0001881883
## bankwest:speciesfranksi 0.0377029180
## difference
## pvalue bankeast:speciesfranksi
## bankeast:speciesfaveolata -0.354101893
## bankeast:speciesfranksi NA
## bankwest:speciesfaveolata 0.003861344
## bankwest:speciesfranksi 0.382935078
## difference
## pvalue bankwest:speciesfaveolata
## bankeast:speciesfaveolata -1.2975229
## bankeast:speciesfranksi -0.9434210
## bankwest:speciesfaveolata NA
## bankwest:speciesfranksi 0.1360547
## difference
## pvalue bankwest:speciesfranksi
## bankeast:speciesfaveolata -0.6926464
## bankeast:speciesfranksi -0.3385445
## bankwest:speciesfaveolata 0.6048764
## bankwest:speciesfranksi NA
##
## $H35JRAZ03DJU1V
## difference
## pvalue bankeast:speciesfaveolata
## bankeast:speciesfaveolata NA
## bankeast:speciesfranksi 0.55695854
## bankwest:speciesfaveolata 0.01260529
## bankwest:speciesfranksi 0.01260529
## difference
## pvalue bankeast:speciesfranksi
## bankeast:speciesfaveolata -0.663134134
## bankeast:speciesfranksi NA
## bankwest:speciesfaveolata 0.003861344
## bankwest:speciesfranksi 0.003861344
## difference
## pvalue bankwest:speciesfaveolata
## bankeast:speciesfaveolata 1.2885484
## bankeast:speciesfranksi 1.9516826
## bankwest:speciesfaveolata NA
## bankwest:speciesfranksi 0.9964772
## difference
## pvalue bankwest:speciesfranksi
## bankeast:speciesfaveolata 1.27154039
## bankeast:speciesfranksi 1.93467453
## bankwest:speciesfaveolata -0.01700804
## bankwest:speciesfranksi NA